A better Google Scholar
Table of Contents
Google Scholar is one of the marvels of the modern science ecosystem. Reportedly run by only a dozen of people and started a decade ago by Alex Verstak and Anurag Acharya, it's the most comprehensive and easier to use search engine there is to find scientific works including non-journal publications like preprints or even personal blogs. Whereas competitors come and go, Scholar remains. But it remains, to some extent unchanged. Sure it has added some features in recent years, described in the team's blog but what would an ideal Google Scholar look lile? This post is about that.
What do we want from a scientific search engine?
There are many ways of using something like Google Scholar: There's finding an answer to a very specific question ("Does cloud seeding work?") to building a general understanding of a field (What Andy Matuschak calls here Syntopic reading) to piddling around in novel ways. These three uses could be broadly ordered in order of concreteness
- Problem solving / question answering (finding answers to relatively narrow questions)
- Knowledge building (by finding an interrelated network of facts that add up to understanding)
- Knowledge generation (by generating new ideas, knowing what was previously known to be unknown or not even that)
These are interrelated: To get an answer to a question may be difficult if the evidence is hard to interpret, and to interpret the evidence one needs to know more about the field. To come up with novel research questions one needs to know what has been done so far, and it helps to know what is being worked on at the moment, as well as what is not being worked on.
Polishing Google Scholar
I asked Twitter how we could improve Scholar, and a few interesting suggestions came through. Interestingly they are are features that are already implemented by another search engine, Scite.ai.
- Filtering search by author (Which scite.ai has)
- Enabling sort by date for all years (Which also scite.ai has)
- Complex boolean queries including those that make use of the citation graph
- Filtering based on the article (or book) type (Also in scite.ai)
- Signposting retracted papers (Also in scite.ai!)
Importantly we'd like to do this without introducing clutter. Scite.ai will show if a paper came from "volume 547, issue 7662, P162-162" of Nature. Virtually no one needs that information. But other than that very minor point, does a better Google Scholar look like... Scite?
Why is it hard to do better?
First, Google Scholar is free. It's hard to compete with free when the free product is pretty good. Google Scholar has top-tier search capabilities. From my experience building a project I'll soon describe, Metasearch, it's very easy to add decent search to a site with the power of Elasticsearch Postgres, but Google takes that to a whole new level. Their results, while not always perfect, are more relevant than those of any other academic search engine I've seen. Search is, after all, Google's core competency. And they keep getting better. Whereas in 2020 searching 'avocado' would return a seemingly niche paper about distributed virtual reality frameworks, in 2021 we get more reasonable results, including a recent article on the production and uses of avocados (I just learned that they can be used for something other than toasts!).
Second, Google's paper database is equally unrivaled. Sure, we all know that terrors and weird stuff lurks in the last pages of any Google Scholar search, but in general if it exists, it is in there: GS is the world's most comprehensive academic search engine (Gusenbauer, 2019) there is.
Third, Google Scholar's interface may not have all the bells and whistles of Microsoft Academic or Semantic Scholar, but the UI is fast. When one is filtering through hundreds of papers at a time, being able to do so quickly is key.
Cost is an easy one to work around: If a product is vastly superior to Scholar then I'd pay for it. But the search capabilities and breadth have to be there, no matter what other features are offered: If I have to second-guess another site and fall back to Scholar to be thorough I may as well just stick to a single site.
My attempt: Meta-Search
An attempt to improve over what Google Scholar, albeit in a narrow way, is a website that I had live for a few months (the code, including data loading from Semantic Scholar, is here) called Meta-Search. The website itself is still up, so you can look at it and check the documentation/about for an idea of how it was set up and its API used.
The idea behind Meta-Search is a simple enough one: When I use Scholar, I find myself searching something, then opening the list of papers that cited that paper, then for each of those, do the same and so forth until there are no more papers to check out. Of course I wouldn't literally open every single one of them! I would use some heuristics to decide what to focus on. This was so that I could find work that contradicts, supports, or adds context to the one I initially found.
A very basic way of doing this without having to use much judgement of what is "a good paper" is to simplify the task to "Given a paper, find every systematic review of meta-analysis that cites the paper". This is what Meta-Search did, with some degree of success.
Building this started in early November 2019, the API was built in a day, then in one more week I had the website up. Adding more features wouldn't have been hard. Replicating Google Scholar itself would be a matter of maybe a month?
Not quite. My site never had the search capabilities of Google Scholar and sometimes the results returned would be a bit nonsensical. I tried to mitigate that using citation counts to sort the results but I bet one could do better with some composite score. And unlike Google Scholar, my site was limited to the data that I could ingest from Semantic Scholar. Which is a lot! Even as of today their dataset is the only publicly available one that has a reasonably large size so I can't really do better without paying to e.g. Elsevier to get access to all of it. But large as it may be, it still falls short of Google Scholar.
Meta-Search was an example of taking a relatively defined workflow that I had a need for, and automating it. But even in this case, it was not as good as manual search, the reason being that Meta-Search would classify something as a meta-analysis or review if it had the words "meta-" or "systematic". But not every review is like that. The right thing to do there is to use NLP to be more clever about whether something is or not a systematic review (which is harder to do if you only have the abstract rather then the full-text).
Semantic Search
Scientific papers are static collections of text and images. There are some exceptions like those hosted at Distill.pub, but even those are dynamic collections of text and images and interactive visualizations.
An scientific paper, when stripped of ancillary information like the literature review or some impact justification, is essentially a series of interlinked claims where the premises are a combination of background knowledge and previous results, novel data presented in the paper, or other claims in the paper. In some cases these arguments will be formal: From the premises the conclusions inevitably follow. This is common in mathematics or computer science. In fields that deal with the real world rather than abstractions1 like biology or engineering those chains will of reasoning will never be airtight, dealing instead with fuzzy relations between fuzzy concepts.
So right now, taken as a unit, the information we have to work with taking the paper as a unit of analysis is: authors, abstract, year of publication, journal of publication, and the citation graph. This is all useful information: authors and journals facilitate finding and following topics of interest, the citation graph facilitates understanding where a paper falls within the rest of the literature and so forth.
This is roughly what Google Scholar gives us and it works well enough.
Keywords
A step up from that are the MeSH categories from Pubmed, a series of tags that are manually assigned to papers by a team of indexers. This is the same as authors adding keywords to their papers, only that the MeSH terms are standardized.
I've personally never used these (nor keywords, unless Google Scholar uses them as part of their search) but you could see how they could be useful in a different setting: I rely on a combination of RSS feeds and strategically placed twitter follows to stay abreast of various literatures. But if I were a researcher in one given field, placing topic-based alerts may make a lot more of sense.
Microsoft Academic as well as Semantic Scholar also use NLP to infer topics from the papers without having to use human indexers. This works okay in some cases, my own essay on Bloom's two sigma appears tagged with "Mastery Learning" or "Psychology" but also "Bloom" and "Sigma." Semantic Scholar tags it with just "Psychology".
But is this useful? As I said, I don't make use of it. And moreover I expect these are not that used (thinking what I would do if I were in academia under a different set of constraints and with narrower interests).
Citation graphs
This is the level of analysis at which Meta-Search worked and one available in one form or another on most platforms. At the very least, one can quickly jump to prior art cited by a paper and also what is harder to do manually: browse later work that cited the one paper that is being looked at. This is very useful and I use it all time! Traversing the citation graph helps get a better sense of the surroundings of a given piece of work, what the seminal work in a given area is, or what work has been built downstream of it. But sometimes this can be inefficient: Consider two papers that are published in the same year in a short span of time, in the same field. They don't cite each other but reading them together can be fruitful. Unfortunately citation-level analysis will not highlight this second paper unless, perhaps a year or two later, a systematic review cites both so by traversal one can go from paper 1 to the review and from there to paper 2.
Google Scholar has a Related papers function and Connected papers adds a visual dimension to the same functionality:
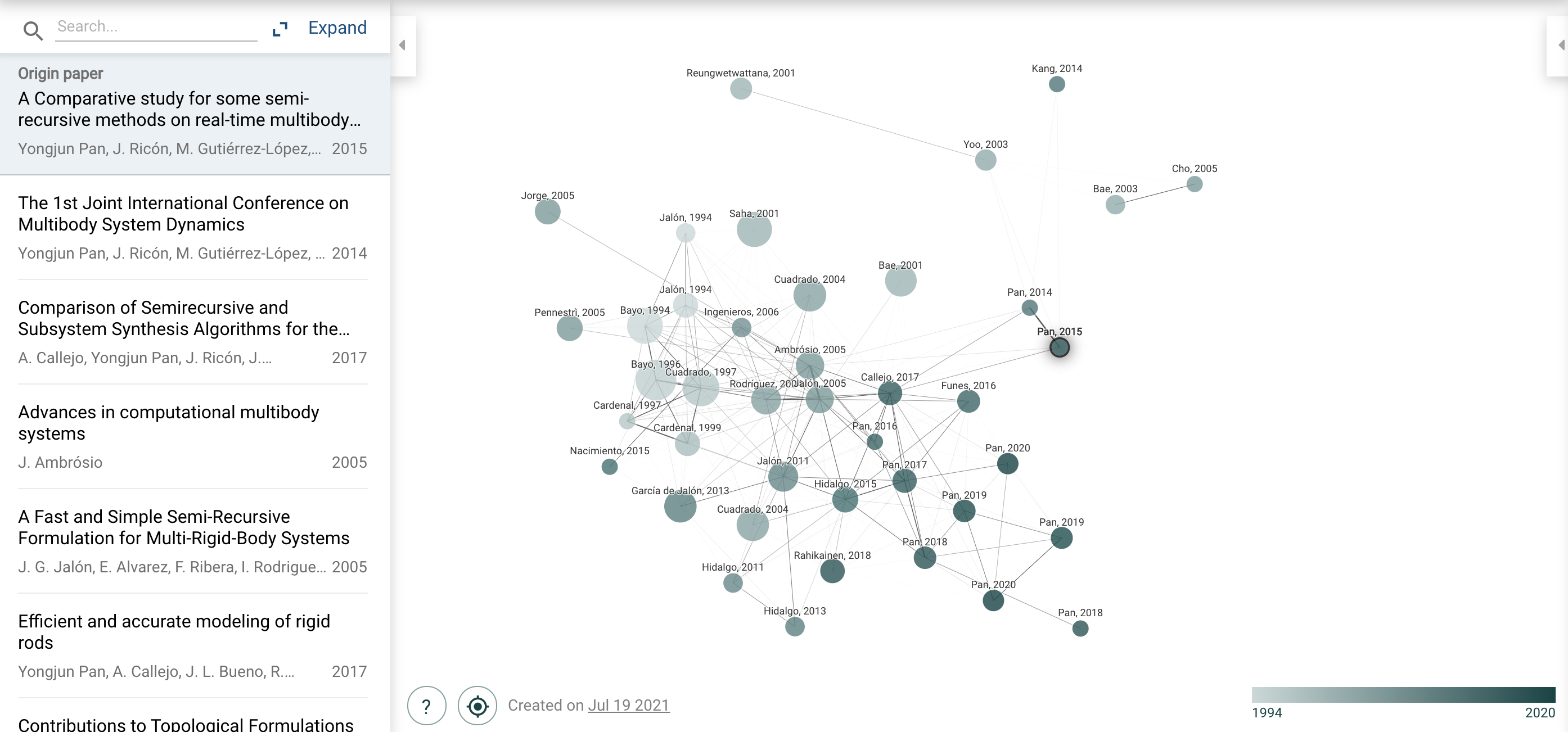
I don't really use either. But maybe I should?
Claim-level analysis
But papers are not monolithic units, as said earlier, they are collections of claims. When a citation is made, it is done to back up some claim, or to add some context. Often it is obvious what the claim might be (the conclusion of the paper) but not always. In my recent wildfires post, near the discussion of the fire deficit, I make a reference to a figure buried in the supplemental materials of a paper I cite. The citation per se (Marlon et al. (2011) ) doesn't say where in the paper is the data I am referencing (I make this explicit in the post by saying where the figure comes from). The worst offenders here are citations to books that contain multiple chapters. Citing the Handbook of the Economics of Innovation without specifying where in it is the source being referenced would make it extremely difficult to find it!
And besides that, citation-level analysis wouldn't tell us about the nature of the citation: Was something cited to criticism it? To highlight a similar result? Just to add context?
The only tool that goes to this level as of today is Scite.ai. As an example take this clinical trial of metformin for diabetes treatment (spoiler: some exercise and weight loss works better than the drug). One of the findings in that paper was the the benefits of exercise were similar in men or women. Scite's smart citations enable us to see later work that found something somewhat different, for example a citing paper had the claim Also, unlike the DPP [the original paper] [...] we found that the intervention effect was stronger in men than in women. Which was flagged as "contrasting" the original work.
Unlike citation analysis, claims are substantially harder to analyze. Scite relies on NLP to tease out the intention of a particular piece of text and this doesn't always work. In the Appendix 1 to this post, check out Example 1 for a case where a contrasting citation is not marked as such. And broadly Appendix 2 for the difficulty of searching through these claims, comparing Scite.ai and another tool called SciFact. As the technology is not there yet (or show me otherwise!) we can't yet use a claim-based explorer as a workhorse scientific search engine.
A workaround is to get scientists to encode in some form of meta-data what the claims may be, but this gets impractical quickly enough:
A claim-level decomposition example
Consider this paper. On its face, we could say that the paper is making the following claims:
- Nicotinamide riboside, candesartan cilexetil, geranylgeranylacetone, and MIF098 do not extend lifespan in mice
- 17-α-estradiol extends lifespan in male mice
The design in each case is similar, so let's take this second one. How does it get justified?
- Feed-the-mice-a-compound experiment
- Dose: 14.4 mg/kg of diet
- Mice strain: UM-HET3
- Control N: 303 male mice, 303 female mice
- Subtreatments (all male mice)
- Early start
- Age started: 16 months
- Intervention N: 156
- Result: 19% increase median lifespan. 7% increase in max lifespan
- Result: 20% weight decrease
- Early start
- Late start
- Age started: 20 months
- Intervention N: 159
- Result: 11% increase median lifespan. 5% increase in max lifespan (not statistically significant)
- Result: 20% weight decrease
- N of sites: 3 (Jackson Laboratory, University of Texas, University of Michigan)
- Statistical method: log-rank, stratification by sites
With a sufficient number of "primitives" we could extract from this relatively simple paper the key results and make them machine-readable for further use. However there is more to the analysis: Lifespan were different across the 3 sites where the experiment was done. Mice were not left to die, rather they were euthanized when technicians at each lab judged that the mice were very sick and about to die soon (Which introduces an element of subjectivity). In this case the diet was standardized, but in another paper part of the same project, results markedly differed by site with the difference attributed to a slightly different diet. Then, we would have to consider diet as well! And suppose that it turns out that the time of the day at which it is administered (or how many times) turns out to be relevant as well, we would have to add that to our list of attributes. Is it realistic to think ahead of time of every possible way in which experiments can differ? I don't think so!
This also highlights an important consideration when looking at claims: that they are fuzzy. The results of this paper, can we take them as "17-a-estradiol extends lifespan in male mice" or do we need to narrow them down to "... in this specific strain of mice" and "...when started at ages 16/20" and "...at this particular dose" or even "... when tested at these particular sites"! A user of this paper could say for example that this compounds extends lifespan in male UM-HET3 mice when started at such ages and be relatively sure of that. But for a paper that takes it further and says that the compound works in mice (in general), this paper would be slightly weaker evidence. Maybe this doesn't work at all in a given strain! Maybe the commonplace C57BL/6J mice requires an altogether different dose. Lastly, if someone wanted to claim "17-a-estradiol extends lifespan in male mammals", this paper would be very weak evidence of that, one would rather need an assortment of similar papers to get there.
And this paper is a relatively simple one, where a compound is fed to mice and lifespan is recorded. Now imagine something slightly complex, say a paper that claims that reviewers have a hard time agreeing on the quality of good research (the Pier 2018 paper I discuss here) and where they conclude that We found no agreement among reviewers in evaluating the same application. Here the key thing the paper is attempting to do hinges upon the link between the word "disagreement" and the way disagreement is measured. As it happens, what we mean by disagreement in common speech is not what the authors mean (Krippendorf's alpha and other measures, which do not necessarily map to the common sense meaning of disagreement).
In the life sciences this is a particularly thorny issue: High level concepts like "immune function" can be operationalized and measured in multiple ways. In this paper from 2016 where a reversal in epigenetic age is claimed, that is based on measurements taken from a particular kind of cell type. Is that reversal of age systemic, happening in all organs? Or take "telomere length"; telomere length is not "a thing", the numerical value one sees on a paper depends on the method used to assess it and the cell types it is measured on, and common measurement techniques can be very noisy, showing great variability for the same sample assessed in multiple labs (Martin-Ruiz et al., 2015).
Social annotation
Beyond, or rather around, the scientific paper there is context that aids in its interpretation. To some extent this could supplement the need for machine-readable papers. Assume for example that a given body of work is largely correct and that there is a limited number of things to watch out for. These can be, and currently routinely are, pointed out by experts in Twitter threads or PubPeer. Additionally platforms like hypothes.is (What powers the comment system in this very website) enables anyone to comment on specific sentences. Fermat's Library also provides an annotation system as seen for example in the Bitcoin whitepaper.
A problem many of these have is that they are fragmented. Wikipedia is the encyclopedia but today there is no "one true place" for discussion to take place in (Or some form of federated comment system that is widely adopted). I have found Twitter for example more useful than Pubpeer for adding context to many publications.
The internet is littered with the corpses of attempts to make science better, tools for thought, and productivity tools of various kinds. This is true here (Especially if we count zombies): Prior to Fermat's there was Beagle, Scrible, RevNote or even Google Sidewiki! Implementing social annotation may be hard (There's even a W3C standard now!), but getting it to be prevalently used and stick around has been so far impossible.
This is because of a system like this benefits from economies of scale: Wikipedia is the defecto "one true encyclopedia" of the internet, and that attracts contributors. If the driving force for adding comments is helping others or telling someone on the internet that they are wrong, the fisking at scale that a centralized commenting platform would enable. No one has kickstarted the virtuous cycle for web annotation yet, let alone scientific paper annotation.
In practice, these systems require either content-provider buy-in (Say Elsevier or eLife) adopting a platform for annotation or users to install some browser extension that enables that capability which adds a layer of complexity: Science publishing is split across multiple entities whereas Wikipedia is a single one. Editing wikipedia doesn't require installing anything, it just works. There was even an attempt at a coalition back in 2017 that aimed to make it so that within three years, annotation can be deployed across much of scholarship. The year is 2021 and we are far from being there.
But these are not the true underlying reasons: eLife after all has hypothes.is support built in but a quick perusal throught their articles shows that this feature is very rarely used: Even this paper co-authored by Jennifer Doudna over seven years ago with over 2000 citations still has just 3 annotations, individually less informative than the average PubPeer comment. A trial at Nature of an "open peer review" system ended because of widespread disinterest in annotations:
Of the 1,369 papers during this period that passed the initial editorial assessment, only 5% of authors agreed to open peer review. Despite healthy online traffic (“5,600 html page views per week and about the same for RSS feeds” [Nature Editors, 2006: n.p.]), these 71 papers received a total of 92 comments, which were heavily concentrated on only 8 papers; 33 papers (46%) received no comments at all, despite the editors’ best efforts to garner them. The editors reported that “it was like ‘pulling teeth’ to obtain any comments”, that “generally the comments were judged to be more valuable editorially than technically”, and that no comment influenced editors on publication decisions (ibid.). As a result, and despite high levels of author satisfaction with the process, Nature discontinued its open peer review system, and has not returned to it. (Skains, 2020)
The same fate was met by PubMed Commons
PubMed Commons has been a valuable experiment in supporting discussion of published scientific literature. The service was first introduced as a pilot project in the fall of 2013 and was reviewed in 2015. Despite low levels of use at that time, NIH decided to extend the effort for another year or two in hopes that participation would increase. Unfortunately, usage has remained minimal, with comments submitted on only 6,000 of the 28 million articles indexed in PubMed.
Why is this? Unlike back in the glorious blogosphere days where being social norms around commenting were very healthy, encouraging both riffing on other's articles as well as directly criticising them, science does not work that way. Leaving useful feedback can mean giving away a good idea for a future paper to a competing lab; criticising a shoddy paper may lead to diminished professional opportunities in the future and retaliation.
Authors responding to Nature’s post-trial survey indicated that they “were reluctant to take part due to fear of scooping and patent applications” (Nature Editors, 2006: n.p.); likewise, discourse participants “might be reluctant to publicly back-up challenges to published data with unpublished data of their own for fear of being scooped, by others or by themselves” (Knoepfler, 2015: 223). This is likely more of a concern in the science, engineering, and design fields, where being first is often paramount to publication or patent grants.
Sometimes building infrastructure can change the underlying values. But here it seems that it's the opposite: the infrastructure is there, but it's just not leveraged. If anything, ironically, there is too much infrastructure/annotation platforms, making it harder for network effects to arise around a single platform.
But if this is so, how come Pubpeer has managed to become the de facto platform for commentary on papers? Perhaps a combination of a) Authors get notified when their paper gets a comment (Which may make it more likely that they engage with the platform) b) Pubpeer allows anonymous comments, alleviating the issues described before c) Pubpeer is moderated, the platform has some guidelines for what sort of comments one can make and perhaps d) Pubpeer very much welcomes the kind of comment that is most likely to be made, the critical one. The fact that their FAQ has a section named "Is someone out to get me?" is telling.
A better Google Scholar
Putting everything together, what would a better Scholar look like? Below is an attempt, using Figma:
This mockup starts by taking the existing Google Scholar and removing the "include citations" and "include patents" which I never use. Sorting has been moved to the top whereas filtering remains in the sidebar. Filtering by date remains there, but now with explicit time bounds by default. One of the Twitter requests was to add filtering by element type (which here I copied from Scite.ai) and I have also added filtering by Systematic review or meta-analysis status as I use that a lot. As with Scite, badges would display Retractions/Errata/Expressions of concern. Pubpeer comments (Or from other sources) are added below the paper, next to the citations. Unlike Scite, I only show the title, the year, the author(s) and the journal.
The functionality provided by Meta-Search would be subsumed by that "Find all downstream work" button, that would fetch everything that cited that paper (and everythin that those works was cited by and so forth), sort it, and return it in a single browsable page.
The largest divergence from the original Scholar is adding a rating system. These stars could either reflect how used the paper is (By clicks on it) or explicitly would enable users to add a rating to any one given paper. The star ratings displayed would include those from academics (say using a registered academic email address) and/or any other visitor to the site.
Not included here is graph-based browsing (That would be its own tab).
This is one way to make Google Scholar better. There may be others!
Appendix 1: Scite.ai
Example 1
An example is a recent paper in the geroscience field that found that a carbon compound (C60) in olive oil, when fed to mice, does not extend their lifespan. The authors of the paper are replying to a small literature that found in the past positive effects from this compound,
Baati T, Bourasset F, Gharbi N, Njim L, Abderrabba M, Kerkeni A, Szwarc H, Moussa F. The prolongation of the lifespan of rats by repeated oral administration of [60]fullerene. Biomaterials [Internet]. Biomaterials; 2012 [cited 2020 Jun 1]; 33: 4936–46. Available from: http://www.ncbi.nlm.nih.gov/pubmed/22498298
Quick KL, Ali SS, Arch R, Xiong C, Wozniak D, Dugan LL. A carboxyfullerene SOD mimetic improves cognition and extends the lifespan of mice. Neurobiol Aging [Internet]. 2008 [cited 2017 Jan 16]; 29: 117–28.
So if I search any of those papers, does the critical paper appear among the citation? The citation is there, but tagged as 'mentioning' rather than 'contrasting'. This is not totally unreasonable on Scite's end: Scite's NLP is sentence-level so if there's a sentence that says 'Our results stand in contrast to those of X' then that will be picked up as contrasting. But in this particular case, there are no such sentences. Rather, the reason that one can know that the paper is clearly contrasting a previous finding is the title of the paper (C60 in olive oil causes light-dependent toxicity and does not extend lifespan in mice) and then fact that it
Example 2
Metformin may or may not extend lifespan in mice. Currently, the highest quality data, coming from the multi-site, large-sample Interventions Testing Program (ITP) at NIA suggest that it does not. A paper they cite that did find effects is the straightforwardly named Metformin improves healthspan and lifespan in mice. We should find the ITP paper shown as contrasting the earlier paper. Once again, the paper that should be "contrasting" the original finding is found as "mentioning", for the same reason as the previous example.
Example 3
I recently came across this paper that successfully replicated earlier work from Cialdini showing that people’s likelihood to comply with a target request increases after having turned down a larger request. In that case Scite correctly found the new paper and tagged it as "supporting", but it did so at a low confidence (56%) and based on a not-so relevant text snippet.
Example 4
Lastly, from my recent wildfires post I found a report that updated the state of the art from where this 2003 report left it. Whereas Google Scholar has over 100 citations for the 2003 report, Scite has 3, none of which is the report. This is probably because the report is technically not a peer-reviewed journal-published paper and Scholar has very lax rules to decide what to index.
Example 5
Another example was this paper from a decade ago, this time in the genetics of longevity. The paper was retracted due to a technical error. The paper correctly shows up as retracted in Scite. This is a feature that's not present in Google Scholar (the paper seemingly is taken down altogether!) or Microsoft Academic, or Semantic Scholar. Only Pubmed (that I could find) also shows retractions.
Appendix 2: Claim-level checking with Scite.ai and SciFact
Scite.ai and the Allen Institute's SciFact aim to provide claim-level analysis based on the full text (in the case of Scite) or the abstract (SciFact). One can search for the claim "The coronavirus cannot thrive in warmer climates." and one gets some results in either platform indicating that the claim is or not supported by various work. The problem is that in both cases, search remains a hard problem and slightly varying the claim leads to different results
- Oxalate causes kidney stones
- SciFact: no results
- Scite: > 5 papers shown as mentioning the claim
- Saturated fat causes cvd
- SciFact: no results
- Scite: > 5 papers shown as mentioning the claim
- The coronavirus cannot thrive in warmer climates.
- SciFact: 2 papers confirming, 2 refuting
- Scite: No results
- The coronavirus cannot thrive in warmer climates (Taking the final period out)
- SciFact: 2 papers refuting, 0 papers confirming
- Scite: no results
- Covid cannot thrive in warmer climates
- SciFact: No results
- Scite: No results
- Zinc reduces flu symptoms
- SciFact: 4 papers supporting
- Scite: 4 papers mentioning the claim
Citation
In academic work, please cite this essay as:
Ricón, José Luis, “A better Google Scholar”, Nintil (2021-07-21), available at https://nintil.com/better-google-scholar/.
Comments