Applied positive meta-science
As scientists we must accept that the world has limited resources. In all fields we must be alert to cost-effectiveness and maximization of efficiency. The major disadvantage in the present system is that it diverts scientists from investigation to feats of grantsmanship. Leo Szilard recognized the problem a quarter-century ago when he wrote that progress in research could be brought to a halt by the total commitment of the time of the research community to writing, reviewing, and supervising a peer review grant system very much like the one currently in force. We are approaching that day. If we are to continue to hope for revolutions in science, the time has come to consider revolutionizing the mechanisms for funding science and scientists.
-Rosalyn S. Yalow, medical physicist (Peer Review and Scientific Revolutions, 1986)
A while back I wrote Better Science: A reader and even earlier "Fixing science (Part 1)". There never was a followup up to those until now. I also once tweeted the diagram below:
Basically everything that has ever been said in the "science is broken" conversation (with some baseline plausibility; disaregarding for now relative importances): pic.twitter.com/IoU7daspDX
— José Luis Ricón Fernández de la Puente (@ArtirKel) May 11, 2020
In this post I initially wanted to think through a categorization of proposals to fix or reform science. There's fixing the replication crisis proposals, various novel funding schemes, new institutions, building tools to help scientists, etc. In thinking about the first axis that becomes salient I immediately jumped to "fixing what's broken" vs "making what's good even better". I tend to be pulled towards the former rather than the latter. My favorite proposal among my New Models for Funding and Organizing Science is after all the Adversarial Research Institute, a novel institution that would not do any new research, but rather strengthen and challenge the existing body of work.
So instead I thought it'd be interesting to explore reforms from a more positive angle as I think most of the discourse around "fixing science" tends to focus on fixing what's broken rather than on improving what is working. Some scattered thoughts on the topic, take this to be working notes to clarify my own thinking, somewhat edited:
Imagine we start tomorrow a research institute in a domain where results are generally known to be reliable. Scientists at this institute will be freed from having to apply for grants, and they will enjoy generous budgets to buy state of the art equipment if they need it. What's more, they will only publish in an open-access free-to-publish publication launched by the institute itself (As Arcadia will do). They can work on whatever they want.
Can we reasonably believe that their productivity can be further improved?
In this example, I've set aside some of the most pressing concerns in applied meta-science: no need to worry about unreliable results, hard to follow methods sections, uncertain funding and job stability, and so forth.
One first family of ideas is to scale up the kind of science that gets done. There are projects that require either coordination between multiple entities, or a large team working on the same goal to produce a new tool or dataset. By setting up an institution that allows for large teams of staff scientists and technicians to execute on a single vision either in one organization (FRO) or across many (an ARPA-type program), we can unlock a different type of work than the one done today at academic labs. This is the driving insight behind Focused Research Organizations and riffs on ARPA like PARPA. This type of organization starts with a goal in mind and works backward to get there. Sometimes the building blocks for the next piece of technology, or the right tools to ask a longstanding question are just lying there waiting for someone to combine them in the right way. Related to FROs there are the related ideas of "architecting discovery" and "bottleneck analysis".
But this all works only if there's a relatively clear path to a goal.
Finding more building blocks
Genetic engineering had been possible for a long time, but there is a qualitative difference between randomly integrating genes into DNA, sometimes by shooting them with a gun into cells, and doing so precisely, in specific sites of the genome. CRISPR gets lots of attention, but before CRISPR, the 2007 Nobel Prize in Physiology of Medicine rewarded Capecchi, Evans, and Smithies for introducing specific gene modifications in mice. This was the first time a targeted edit was made in vivo. The method they used relies on flanking the sequence to be inserted by DNA that matches the sequences that flank the target side. If this sounds similar to guide RNAs is because the idea is exactly the same: using unique DNA (Or RNA, or even proteins) sequences to identify specific unique sites in the entire genome, guiding a variety of effector proteins to the desired site. Conceptually, all gene editing tools we have, CRISPR, TALENs, ZFNs, meganucleases, find their way to the target spot in the same way. Many of these were found from bacteria, not engineered ex nihilo. Hence prior to CRISPR, if one had tried to think of what comes next, the idea of using some sort of sequence-matching guide attached to some protein, and lifting this out of some bacteria would have been the obvious guess. This is exactly what happened, but it would not have been possible without prior discoveries that were wholly unmotivated by gene editing. But "guide+effector protein+desired edit" is one paradigm. It may be the only one we have right now but it doesn't have to be the only one. I don't have much idea of what other ways of making precise gene edits would be, but reprogramming shows an alternative way to approach this question. If asked to roll back the state of a cell to what it originally was (a stem cell), the within-paradigm answer is finding 7000 guide RNAs or similar, and making a few dozen thousand edits in the epigenome until it matches what it used to be. But it is much easier than that: delivering four transcription factors (the "Yamanaka factors") works. The out-of-paradigm way of thinking is not specific genes, but cell state in general. Admittedly this is cheating because the machinery to reprogram the cells is already in the cells, and we don't know yet how reprogramming works, but it's a proof of concept of cell state engineering (as opposed to genetic engineering).
Molecular recording (for example Linghu et al. 2021) is less famous than CRISPR, not very widely used but very interesting nonetheless. Molecular recording is getting cells to record what they do in molecular ticker tapes, as it happens in real time. You could imagine a neuron recording its history of spike activity in a very long protein ticket tape. Then you could imagine all neurons doing this. At that point you have a full recording of what a mammalian brain has been doing for a few minutes. We're not there yet but we're at the point where we can make cute cell pictures with the ticker tapes inside of them:
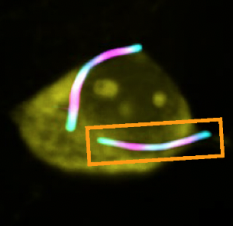
How do you make this? In this case, first you want something that can form filaments inside cells. The Linghu paper says 14 of these proteins were tested, they picked one and ran with it to develop it into a tool capable of making cells happily record themselves. Why those 14? Probably someone searched Google Scholar and found some suitable candidates characterized in prior work. Now one could imagine taking those 14, doing some ML to generate more candidates, and then trying to explore the space of possible proteins to find others that are even better, but the point is that in this case a very deliberate invention relied on a plethora of unrelated discoveries, just as with CRISPR. In the ideal scenario the input "Give me proteins that can form filaments and get fluorescent tags attached to them" wouldn't only output what we know, but something like "Spend 10% of your resources in randomly sampling the Earth for new microbes, 40% poking at this one specific genus of bacteria and 50% on running AlphaProtein1 . This seems to require full-fledged AGI.
If not for the Mojicas of the world we wouldn't have the building blocks needed for building new tools: might we have to conclude that, as one book put it, greatness cannot be planned? That we need undirected explanation to find building blocks?
I have to admit, the title of this book bothers me a lot. The point of the book is that on the way to get to X, exploration of unrelated areas without the objective of getting to X in mind can lead to key stepping stones to get to X. So to pursue X it's best to spend some resources not pursuing X at all.
To an engineer-minded person like me, serendipitous discovery sound like a sadly inefficient affair. It's something (as with tacit knowledge) to be acknowledged as real, but not celebrated. It's giving up on the dream of an assembly line of knowledge, accepting a somewhat romantic view of science where we have to give up on reasoning about the best path forward from first principles2 .
Prime Editing (Anzalone et al., 2019), a recent and more accurate successor to the original CRISPR was developed with an objective in mind (more accurate gene editing), it was not found by chance. But its building blocks were: One of the components of Prime Editor 1 (PE1) is a reverse transcriptase enzyme from the Moloney Murine Leukemia Virus (M-MLV RT). Who would have told us that a nasty cancer-inducing virus would one day be used for gene editing! They then constructed a bunch of alternate versions by introducing mutations to M-MLV RT. Why M-MLV? Maybe because it's commonly available. Perhaps in an alternative universe, Prime Editing would be powered by avian myeloblastosis virus RT instead.
From these and other examples it seems that we could do a bit more planning for greatness: The process that led to these tools was
- A preexisting library of characterized components
- An idea of how a kind of component would be used for a given purpose
- Trial and error
Step 3, technically known as fuck around to find out exists there where simulation is imperfect. If there are 3-4 candidate solutions and testing them is quicker and/or cheaper than trying to simulate or otherwise prove the optimality of one, then testing them all, then picking the best candidate makes sense. Given that it's hard to simulate biology in silico and a common unit of simulation (cells) are small, the idea of screening libraries of compounds in vitro is pervasive in the life sciences. It's something that works reasonably well and is cheaper and faster at the moment than running a cell on AWS (which we can't do yet).
Step 2 is about generating a good research question, here "How do we get cells to record the expression of one protein over time" which may have been originally motivated by a higher level question "How do we record the entire brain". It is also about having the right knowledge to know what building blocks make sense, and the knowledge of where to go find them.
But if optimization is what we want (Once the properties of the building blocks are known) then instead of relying on the next Mojica to notice weird patterns in genomes, we could be able to optimize or synthesize directly the protein that is needed. In a recent paper (Wei et al., 2021) train a convolutional neural network to predict the performance of guide RNAs. They provide a tool as well to run predictions of what the best guide RNAs are to target given genes. The same applies to Step 1: If we have a large dataset of biological entities and some readouts for what they do, eventually we'll be able to just ask a model to give us what we want. The space of possible proteins that are useful is finite, and relying on what happens to work in nature is but a step towards progressively better tools.
Working towards such models and datasets is one way to plan for, and accelerate, future great discoveries.
Tools for (scientific) thought
But we're far from a world where the best possible gene editing tool comes out of an ML model; that probably comes close to the kind of problem one probably needs AGI for.
Other kinds of software could be useful. Imagine for a moment that scientific search engines like Google Scholar or Semantic Scholar disappear. Imagine going back to manually reading hundreds of volumes of multiple journals to find what you want. Is there a tool that makes our current situation analogous to the pre-Scholar days? Some months ago I tried to riff on Scholar in my Better Google Scholar post, concluding that it's very hard indeed. In practice, the kind of software that one sees in science is not particularly glitzy. Some of it is for lab logistics (Quartzy, Softmouse, FreezerPro), lab notebooks (Benchling), or reference management (Mendeley, Zotero). There are some special purpose software tools to solve concrete problems. BLAST can take sequences of DNA or proteins and return similar ones in other species. Primer-BLAST can generate primers for PCR. GraphPad is commonly used for biostatistics and Biorender for illustrations. And then there's Excel.
It seems to me some of these tools are not like the others. Some tools make life easier, others enable new kinds of experiments. Suppose we go travel back in time and give Crick & Watson Google Scholar and GraphPad. I don't think much would have changed. Give them the tooling used by modern X-ray crystallography and we would have had the structure of DNA faster than we actually did. Similarly, give TPUs to the early pioneers of artificial neural networks and we may have jumped to the present situation without a few AI winters inbetween.
But these tools still are only useful once you have the right question to ask. Or do they? By having more tools, we are able to piddle around in novel ways3 . This piddling can then lead to better questions. For example, the question "How do neural networks learn or represent knowledge" is pointless until we have the hardware to run large models and the visualization tools to look into them. Only at that point can Chris Olah and friends go and wonder and wander.
Perhaps we have to reconsider the idea that "time-saving" tools do something minor. The one lever we know for sure we have to generate new interesting ideas is having more time. So the more time that can be spent thinking and less doing mechanical work like finding a paper, talking to a sales rep to buy a new instrument, or debugging a new protocol, the more interesting thoughts will be thought.
Time is all you need?
At first, I thought that "marginal improvements" that just save time wouldn't be that interesting to think about. Doing the same thing but faster or cheaper doesn't seem like how one gets to interesting new thoughts. But if those thoughts are a function of primarily time spent thinking, then buying time is how one gets more of thoughts: scientists spend a lot of time not only applying for grants, but also pipetting, cleaning lab equipment, and moving samples in our out of instruments. Spending more time reading papers, or doing analysis seems like a better use of their time. Whether we can do better than buying more time is something that we could figure out by surveys of how specific discoveries came to be, for example taking Nobel-winning research, or by talking to scientists directly.
In writing this, I am more confident that at least for me the most useful thing (not the most useful thing period, that's hard to say; rather the thing at the intersection of what I enjoy doing and what I can do) is not to find ways to help us think better, but to take existing building blocks and designing FROs for concrete problems we know we have.
Appendix: A collection of various proposals to improve science
When I started writing this post I tried to think of some high level categories for all the "fix science" proposals. Some of these are discussed in earlier posts, see Science Funding.
-
Software
- Search tools
- Social annotation
- Reference managers
- ML models for replication
- ML for idea generation
-
Tooling
- Cloud labs (Strateos, ECL)
- Lab automation
- Spatial creativity (Dynamicland? Interior design/architecture to foster collaboration & serendipity, MIT Building 20)
-
New institutions
- PARPA
- FROs
- Prediction markets for replication
- Uber, but for technicians
- New journals
-
Funding mechanisms
- Lotteries
- Find the best, fund them
- Fast Grants
- Fund People, not Projects
- Alternative evaluation practices (deemphasise impact factor etc)
- Funding for younger/first time Pis
- Funding high disagreement research proposals
-
Norms
- Preregistration
- Replication
- Fund the right studies
- Mandatory retirement
- Cap on funding
- Better methods sections
- Open Science
-
Activities, norms
- Roadmapping
- Workshops
- Grants to strengthen studies
- Record videos
- Reviews
- Fighting data forging
Citation
In academic work, please cite this essay as:
Ricón, José Luis, “Applied positive meta-science”, Nintil (2022-04-20), available at https://nintil.com/metascience-categories/.
Comments